Earlier in this series, I wrote about why bio/medical AI repositories need more than benchmarks, what I learned after auditing 10 public repositories, and why an AI auditor itself needs a memory contract.
That work led to STEM-AI v1.1.2 and the MICA layer: a memory-contracted initialization step that forces the auditor to load bounded rules before scoring begins.
For the broader arc:
But after that, a different engineering problem took over.
The audit logic was stricter.
The reports were richer.
The reasoning was more bounded.
But the developer workflow still felt too loose.
So the next question was no longer:
How do I score trust?
It became:
How does a bio-AI audit tool become something an engineer can actually run, gate, inspect, and integrate?
The answer turned out to be less about seeing more signals and more about refusing to confuse them.
That is the core argument of this post:
A detector becomes more trustworthy when it is strict about what it cannot conclude.
Once I took that seriously, STEM BIO-AI stopped looking like “one score plus some extra metadata” and started looking like a system with distinct lanes, distinct boundaries, and distinct operator workflows.
The problem was no longer scoring
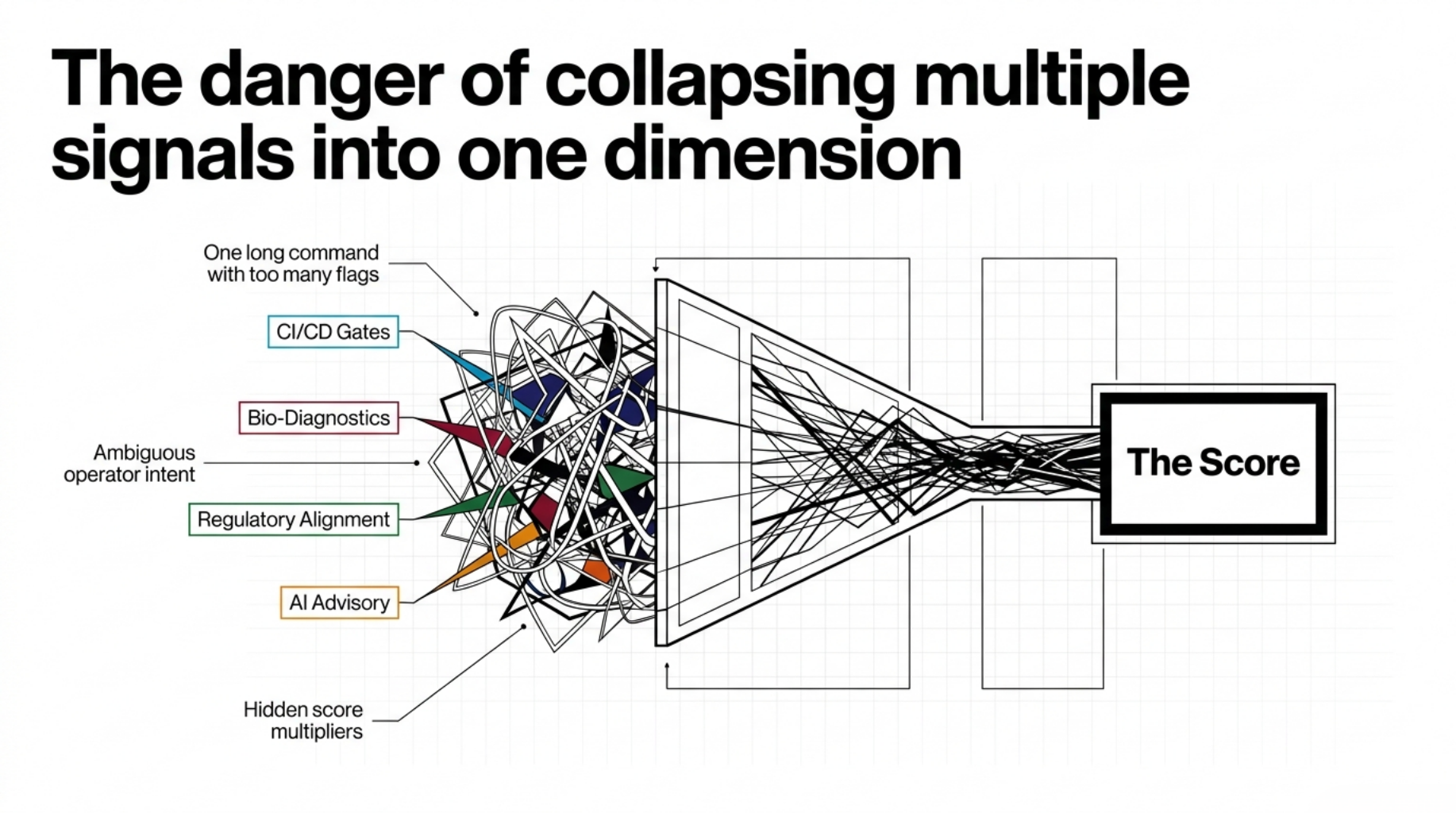
By the time I reached the 1.6.x line, the rubric was no longer the main bottleneck.
The bottleneck was operational clarity.
A trust audit tool is not very useful if:
- the normal path is one long command with too many flags
- CI has to reverse-engineer the result from human-readable stdout
- bio-specific diagnostics are mixed directly into the same surface as formal scoring
- regulatory relevance shows up as vague implication instead of explicit traceability
- advisory AI is present, but its relationship to the official score is unclear
At that point, the tool stops being hard to trust for conceptual reasons and starts being hard to trust for operational reasons.
That is a different class of problem.
The CLI had to reflect operator intent
The earlier CLI was functional, but too flat.
You could do things like this:
stem /path/to/repo --level 3 --format all --explain
stem /path/to/repo --tier-gate T3 --format json --quiet
stem /path/to/repo --advisory packet
stem /path/to/repo --advisory-response provider_advisory.json
All of that worked.
The issue was that it treated very different operator intents as one long option surface.
In practice, these are separate workflows:
- scan a repository and generate artifacts
- enforce a gate in CI/CD
- export a bounded advisory packet
- validate a downstream provider response
- cross an explicit provider-call boundary
So I refactored the CLI around workflows instead of flag accumulation:
stem scan <folder>
stem gate <folder> --min-tier T2
stem advisory validate <folder>
stem advisory packet <folder>
stem advisory call <folder>
stem advisory check-response <folder> --response FILE
The older paths still exist for compatibility:
stem <folder>
stem audit <folder>
stem <folder> --tier-gate T2 --quiet
stem <folder> --advisory packet
But they are no longer the conceptual center.
That matters more than it sounds.
Once the command names match the operator’s intent, the system becomes easier to teach, easier to remember, and easier to wire into pipelines.
This is not just a DX cleanup. In a medical or bio-adjacent audit context, command ambiguity is part of the trust problem.
Repository trust needed four separate lanes
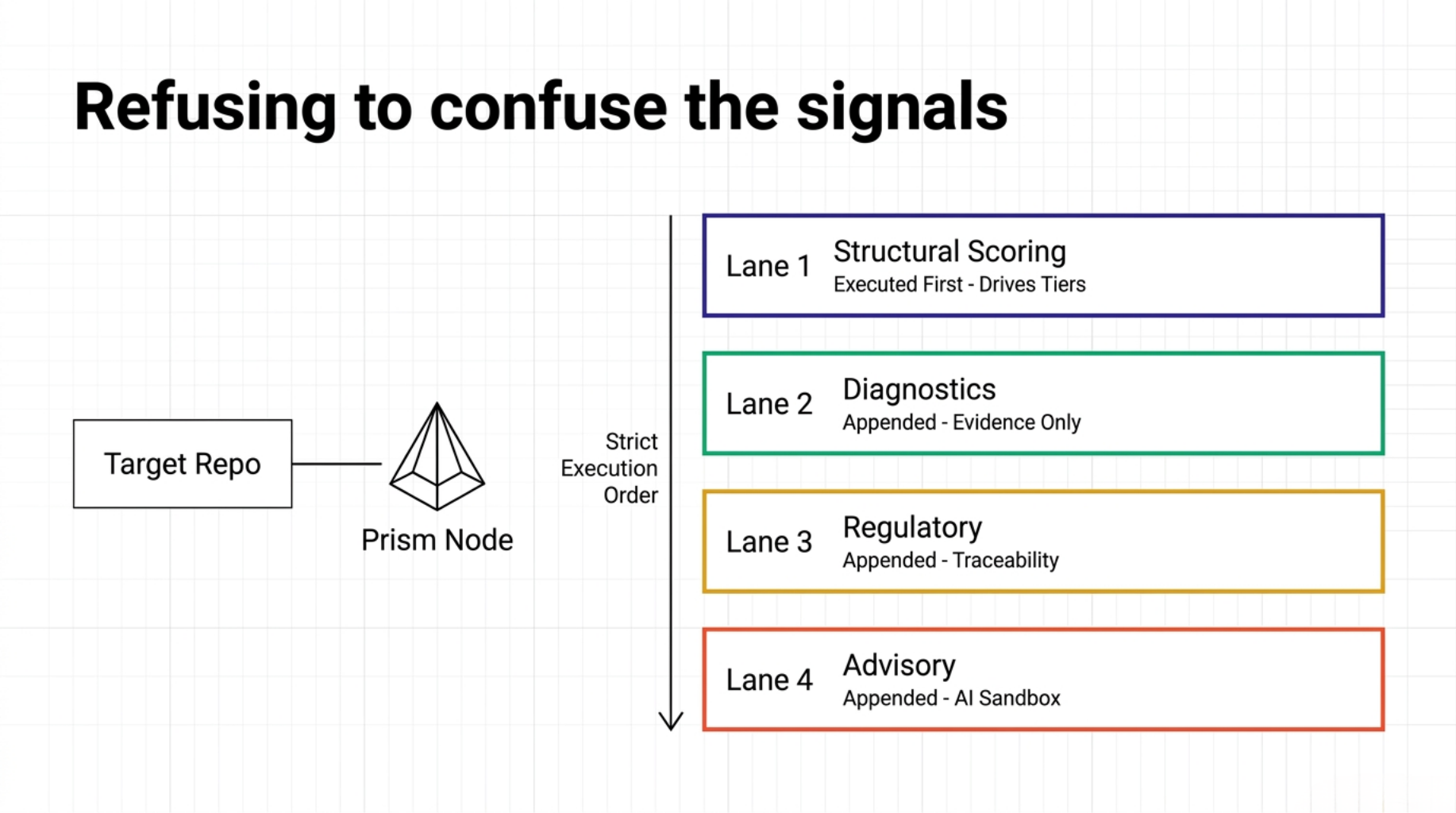
This was the biggest architectural shift.
I stopped treating repository trust as one object.
In practice, it needed four separate lanes:
- deterministic structural scoring
- deterministic diagnostics
- regulatory traceability
- optional AI advisory
If all of those collapse into one final confidence score, the tool becomes harder to reason about.
The more regulated the domain, the more dangerous it becomes to collapse every useful signal into one score.
Some evidence should change the score.
Some evidence should only raise review priority.
Some evidence should support traceability.
Some evidence should be handed to a human or advisory system.
The maturity of the tool is not that it sees all of them.
The maturity is that it does not confuse them.
This separation is not just conceptual. It exists in the code path.
One reasonable objection to any architecture write-up is: are these really separate lanes, or are they just different labels on the same output object?
In STEM BIO-AI, the answer is visible in the execution order.
The scanner computes the formal score first. In the result object, that means keys like:
- Stage 1
- Stage 2R
- Stage 3
- risk penalty
- score cap
final_scoreformal_tier
Only after that does it append the non-scoring layers, again as explicit result keys:
regulatory_basisstage_traceabilityregulatory_traceabilityreasoning_model- optional
ai_advisory
That ordering matters.
The score is not derived from the advisory lane.
The regulatory mapping does not mutate the formal tier.
The diagnostics lane can emit evidence without becoming a hidden score multiplier.
That is why the next four sections exist.
1. Deterministic structural scoring
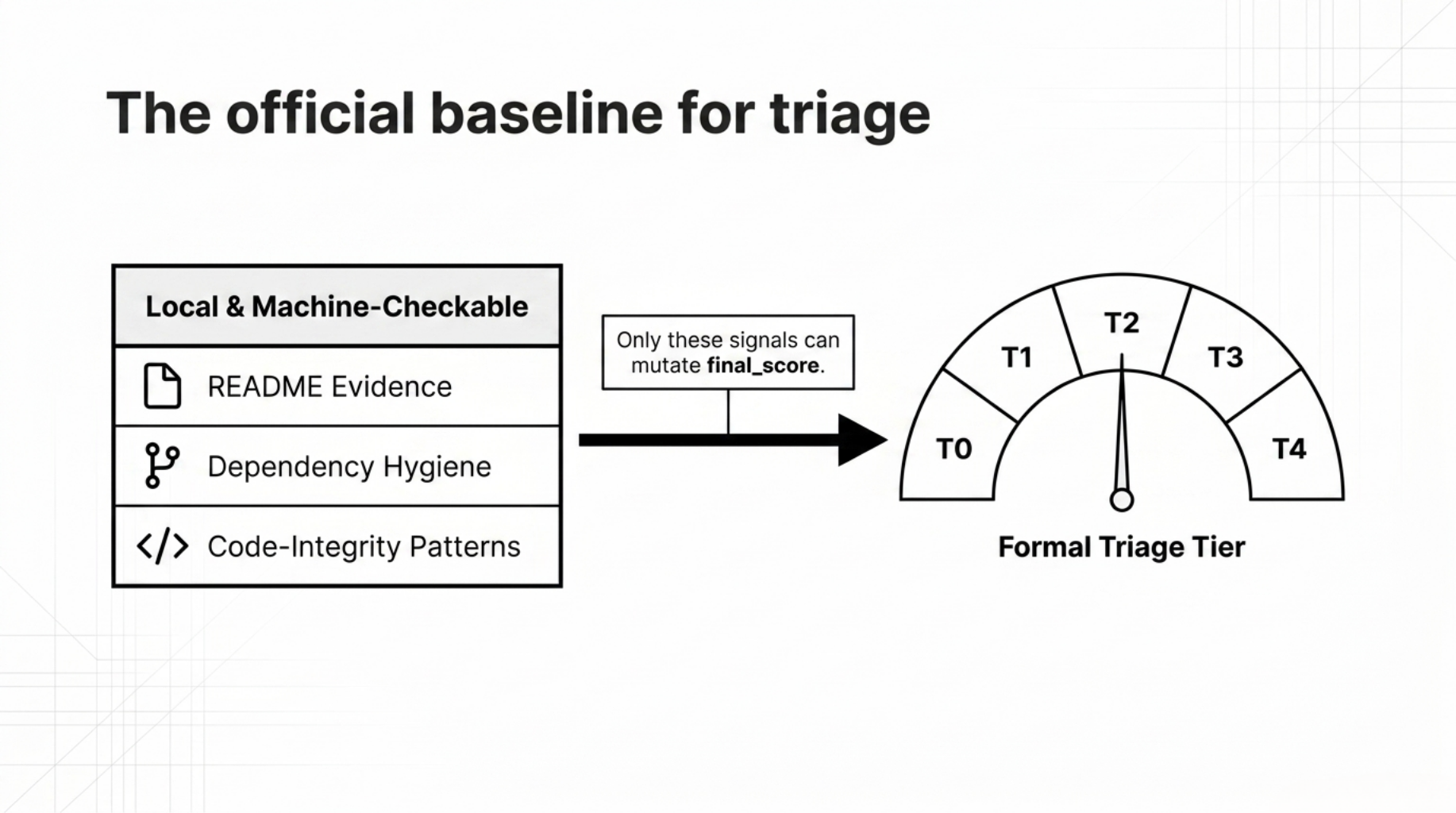
This remains the official score and tier.
It measures the main repository-visible signals:
- README evidence
- repo-local consistency
- code and bio responsibility
- dependency hygiene
- changelog and provenance surfaces
- code-integrity patterns
This lane is local, deterministic, and machine-checkable.
That is the part that can legitimately drive a formal triage tier.
I am not claiming this is the only possible architecture. A different system could have folded diagnostics or replication more aggressively into one unified score.
I chose not to, because the narrower score proved easier to defend. A smaller claim with cleaner boundaries was more valuable here than a broader score with ambiguous semantics.
2. Deterministic diagnostics
This is where the deterministic diagnostics spec became important.
I needed a place for findings that are real, useful, and inspectable, but should not silently perturb the main score until they are calibrated.
That is what docs/DETERMINISTIC_DIAGNOSTICS.md defines.
It separates the diagnostic problem into two lanes:
- Lane A: deterministic local diagnostics
- Lane B: optional AI-assisted semantic review
That separation is central.
The deterministic lane is authoritative for hard findings.
The AI lane is advisory only.
The local diagnostic lane currently focuses on evidence-bearing bio-specific signals such as:
- malformed or suspicious SMILES-like outputs
- missing parser guards
- silent mock or simulated-data fallbacks
- risky subprocess construction around bio tools
- traceability manifest surfaces
The point was not to create a “bio slop detector” with a catchy label.
The point was to create a local evidence lane that could say:
- here is the file
- here is the line
- here is the snippet
- here is the bounded interpretation
That is much more useful than a vague semantic warning.
Why diagnostics stayed evidence-only
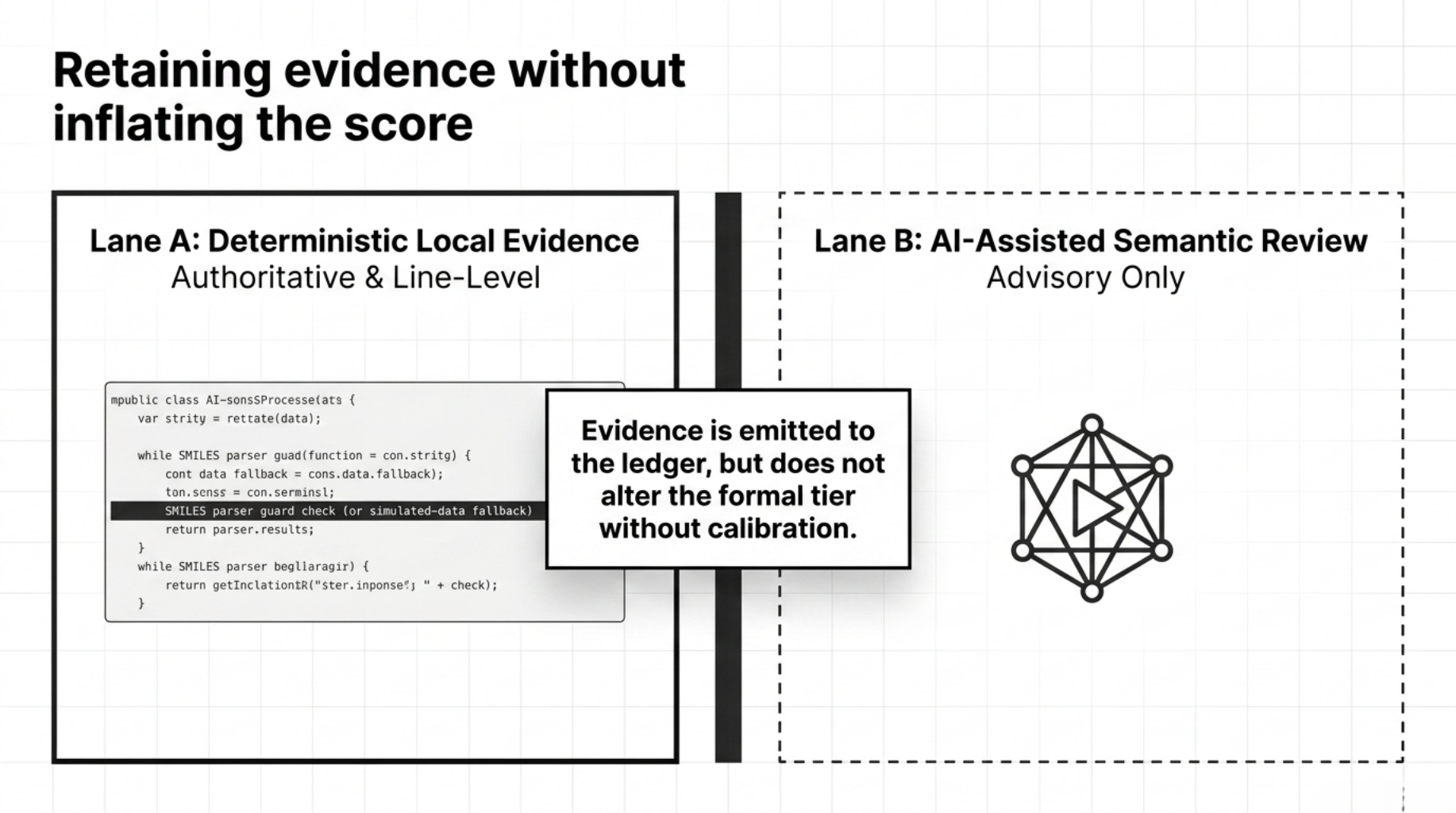
This was one of the harder engineering decisions.
It would have been easy to push every new bio-specific detector directly into the final score.
I did not do that.
The deterministic diagnostics spec is explicit that many of these findings begin as evidence-only. In practice, they are emitted as line-level records in the result object's evidence_ledger:
- findings are emitted into the result object’s
evidence_ledger
- findings appear in Markdown and
--explain
- findings do not change
final_score or formal_tier
That is the right default.
For example, the SMILES lane can be very useful for detecting:
- malformed surface strings
- low-entropy placeholders
- repeated trivial outputs
- missing parser guards
But it does not prove:
- medicinal usefulness
- synthetic feasibility
- binding plausibility
- biological efficacy
- full chemical validity in every edge case
That boundary is important.
A detector becomes more trustworthy when it is strict about what it cannot conclude.
Evidence-only is the temporary safe default until a detector has earned score authority through commit-pinned benchmark evidence, false-positive review, and reproducible calibration.
3. Regulatory traceability
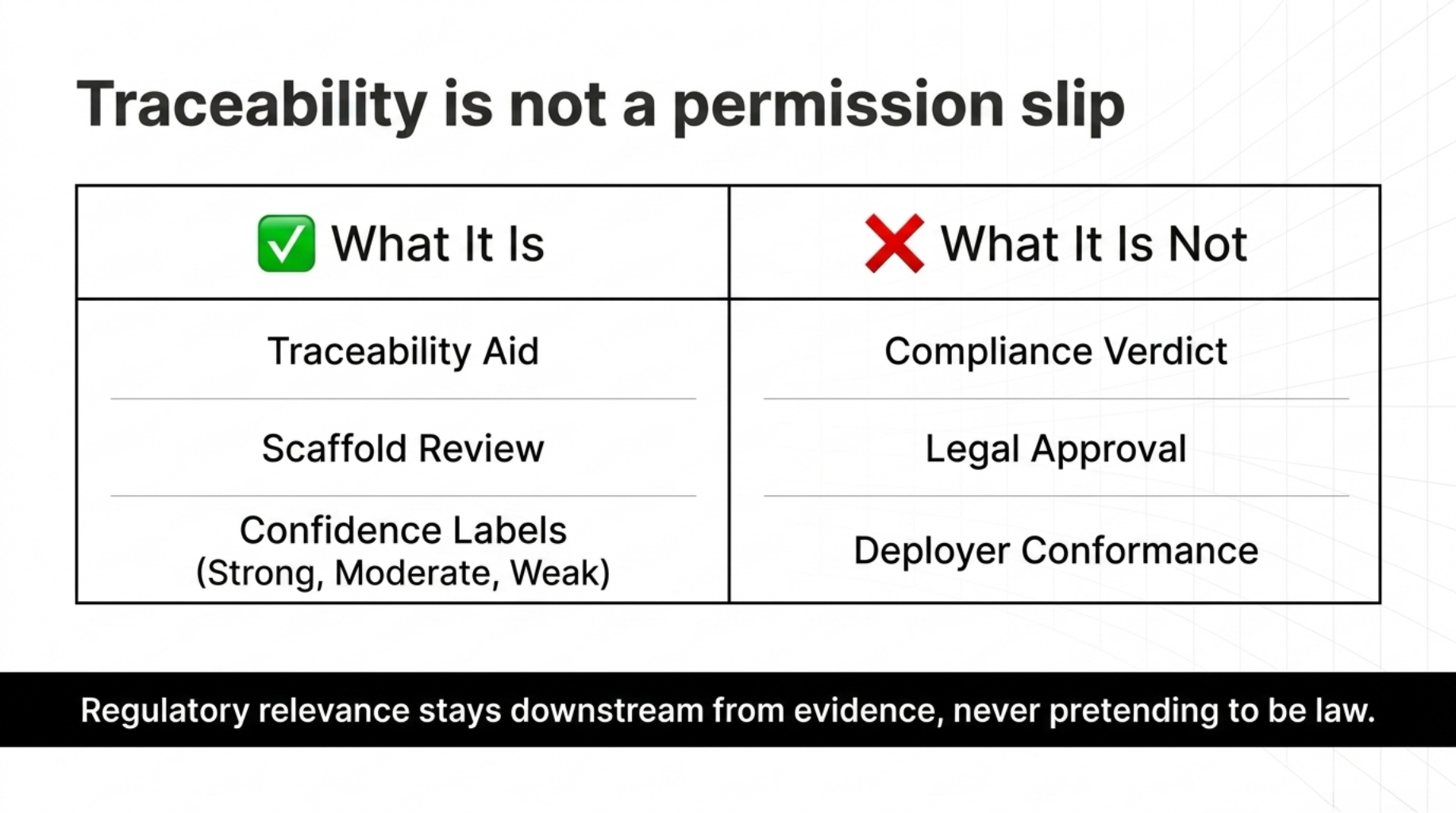

The second document that became central was docs/REGULATORY_MAPPING.md.
This solved a different problem.
Once you audit clinical-adjacent repositories, people naturally ask:
- does this align with EU AI Act themes?
- does this help with FDA-oriented review?
- is there anything relevant to IMDRF or SaMD evidence families?
The wrong answer would be to turn those questions into a fake compliance score.
So I did the opposite.
The regulatory layer is explicitly framed as:
a traceability aid, not a compliance verdict
That document maps observed evidence classes to requirement families with bounded confidence labels like:
- strong
- moderate
- weak-moderate
- weak
- not assessed
the confidence applies to the mapping relationship, not to legal acceptability.
That means the tool can say things like:
- versioned manifests and changelogs may support record-keeping / traceability review
- intended-use and disclaimer sections may support transparency scaffolding review
- override interfaces may support human-oversight interface review
- subgroup measurement language may support weak evidence of data-governance intent
without claiming:
- legal compliance
- regulatory clearance
- clinical certification
- deployer conformance
In a regulated domain, traceability is useful only when it does not pretend to be permission.
A concrete example: why Article 12 is traceability, not compliance
The best example here is EU AI Act Article 12 style traceability.
The regulatory mapping layer treats signals like:
The regulatory mapping layer treats signals like changelogs, checksum manifests, versioned config surfaces, audit-log schema fragments, and decision-event or override-event schema tokens as evidence that a repository may have traceability scaffolding.
That is useful.
It is also bounded.
Changelog presence is not the same thing as deploy-time event logging. Current scope does not establish runtime log completeness.
So the output can legitimately say:
there is structural evidence relevant to traceability review
while refusing to say:
this system satisfies traceability obligations
What this buys in practice is not a compliance shortcut, but a faster review question. If a repository exposes none of the scaffolding signals in this lane, there is little reason to treat it as traceability-ready for deeper institutional review. If those signals do exist, the next step is still expert inspection, but the scanner has opened the right folder and pointed at the right files.
Why regulatory mapping stayed subordinate to evidence
Regulatory relevance had to remain downstream from evidence, not a score multiplier pretending to be law.
That is why the output shape separates things like:
regulatory_basisstage_traceabilityregulatory_traceability
from the actual score computation.
The regulatory basis object is registry-driven. It can mark review_required when the basis registry is stale or required source families are missing. That is a traceability control on the mapping layer itself, not an input into the scoring formula.
If a repo has traceability-relevant scaffolding, that is useful.
If a repo has traceability-relevant scaffolding, that is still not compliance.
The distinction has to remain visible in both the code and the artifacts.
4. Optional AI advisory
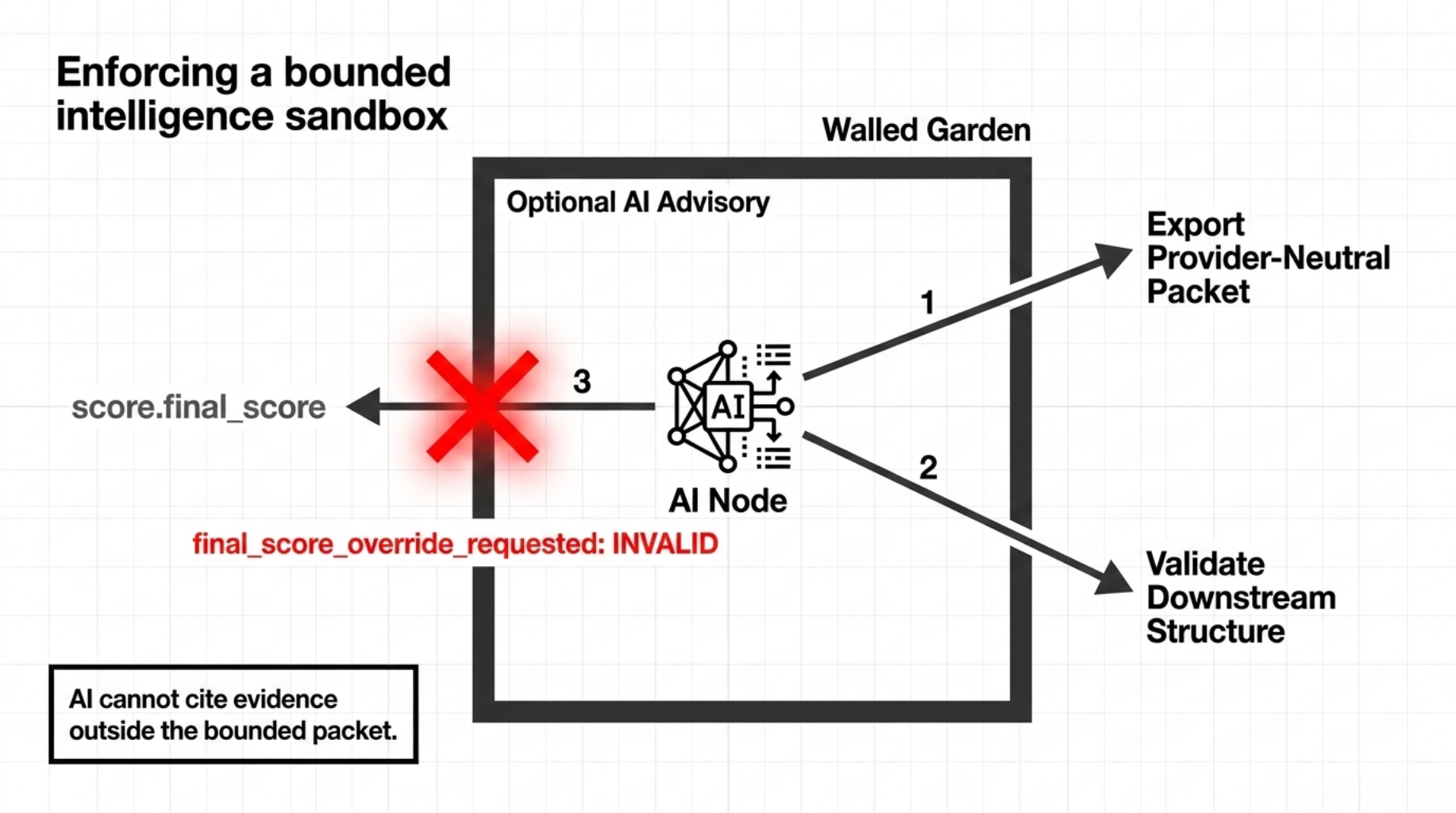
The fourth lane is the advisory layer.
This one exists for bounded model-assisted review, but it does not get to rewrite the official outcome.
That means workflows like:
stem advisory packet /path/to/repo
stem advisory check-response /path/to/repo --response provider_advisory.json
can exist without creating ambiguity about who owns the formal result.
The advisory layer can:
- export a provider-neutral packet
- validate downstream response structure
- enforce finding-ID citation rules
- reject prohibited claims
- surface runtime and secret boundaries
What it cannot do is silently override:
score.final_scorescore.formal_tier
How that rule is actually enforced
The advisory validator explicitly checks for score-override attempts. If a response includes fields like:
final_scoreformal_tierreplication_scorereplication_tier
or sets final_score_override, the response is marked invalid with final_score_override_requested.
The packet contract also exports the rule in plain language:
Do not modify or override final_score, formal_tier, replication_score, or replication_tier.
Provider responses must cite exact values from allowed_finding_ids; citation strings are not repaired or loosely matched later.
So the advisory lane is bounded in two ways:
- it has no authority to change the deterministic result
- it cannot cite evidence outside the bounded packet
That is the kind of mechanism I mean when I say “better boundaries.” If the rule cannot be checked, it is not really part of the architecture yet.
What operational use looks like now

Once these lanes were separated, the CLI became much easier to reason about.
Local engineering review:
stem scan /path/to/repo --level 3 --format all --explain
CI/CD gate:
stem gate /path/to/repo --min-tier T2 --summary off --output results
Offline advisory packet generation:
stem advisory packet /path/to/repo --output advisory_out
Downstream provider response validation:
stem advisory check-response /path/to/repo --response provider_advisory.json
The important point is not just that these commands exist.
It is that each one represents a distinct trust boundary.
That made the project feel more like engineering infrastructure and less like a scoring demo.
A real v1.6.2 packet
To make that less abstract, I re-ran STEM BIO-AI v1.6.2 against a local clone of ClawBio, which describes itself as a local-first, privacy-focused, reproducible bioinformatics-native AI skill library.
The command was:
python -m stem_ai.cli scan /path/to/ClawBio --level 3 --format all --explain
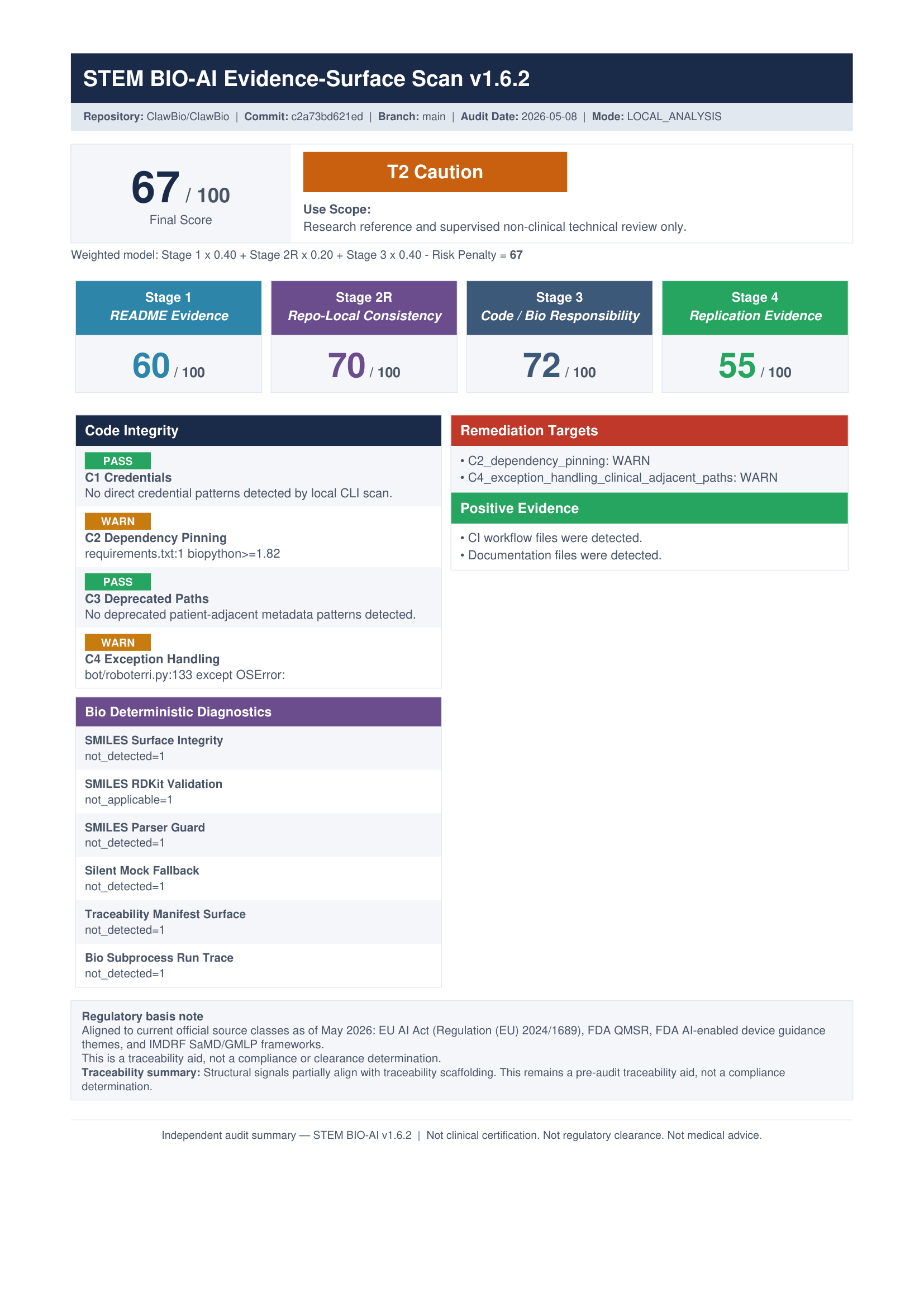
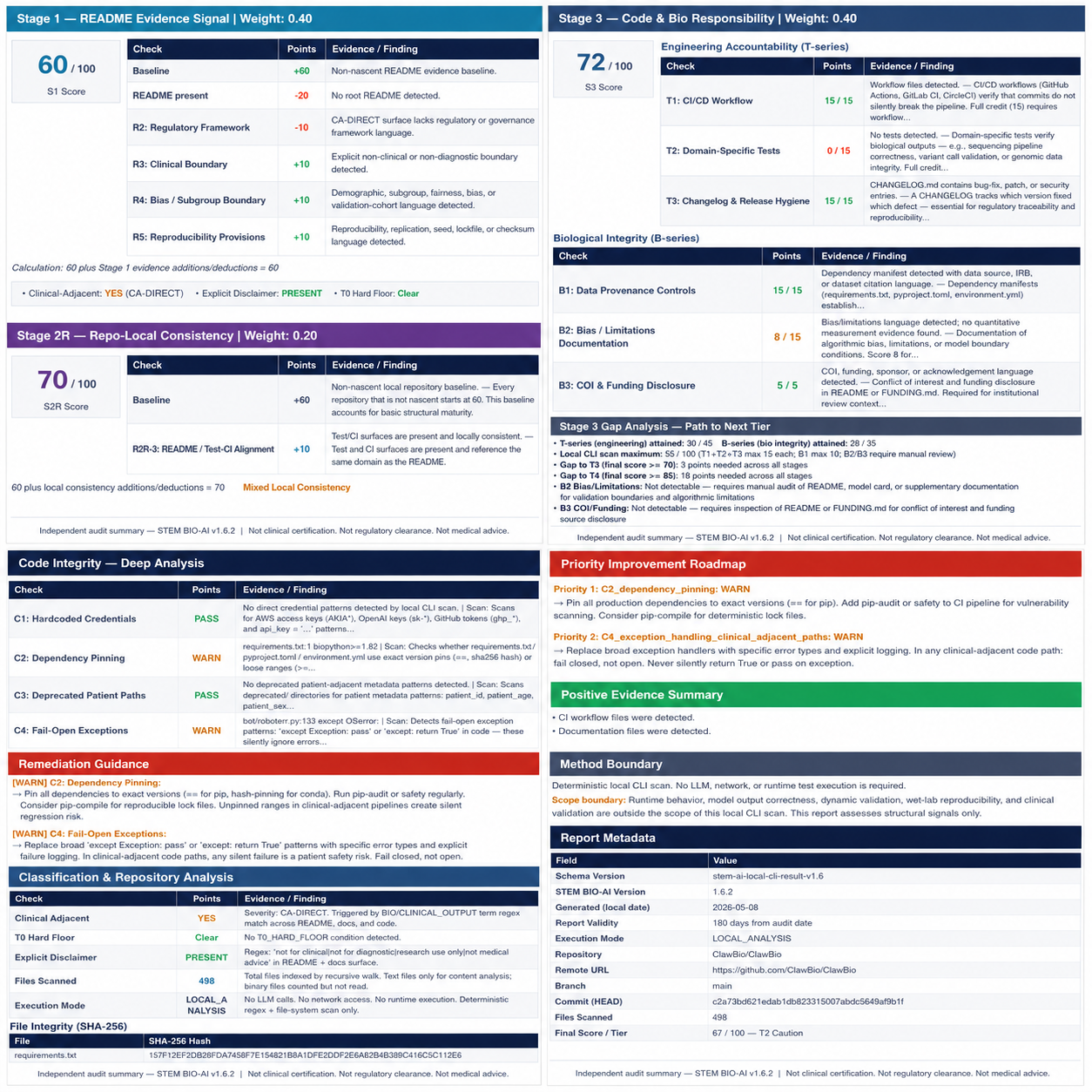
On my machine, the scan took about 9.4 seconds and produced the normal artifact set:
JSON for automation
Markdown for review
PDF for human-facing packet inspection
--explain for file / line / snippet proof tracing
The result was:
- 67 / 100
- T2 Caution
- Replication lane: 55 / 100 (R2)
- Clinical adjacency: CA-DIRECT
- Code integrity warnings: C2 dependency pinning, C4 exception handling
Two details mattered.
First, the scanner did not manufacture chemistry findings just because the repository was bio-adjacent. The SMILES diagnostics stayed not_detected or not_applicable.
Second, the scanner enforced observable repository conventions. Because ClawBio used ClawBio_README_Repo.md rather than a root README.md, the scan recorded S1_missing_readme: -20.
A human reviewer might contextualize that differently. The scanner does not. It records what the repository exposes through the surfaces it knows how to measure.
That is the point of the workflow: not to replace review, but to make the review surface inspectable.
What still has to stay bounded
The system is better than it was, but three boundaries still matter.
First, the public surface is broad. Scoring, diagnostics, replication, advisory packets, regulatory traceability, JSON, Markdown, PDF, and explain traces are all useful, but they increase onboarding cost.
Second, diagnostics should stay evidence-first until they have benchmark-backed calibration. Commit-pinned fixtures, reproducible output, and explicit false-positive review should come before score authority.
Third, regulatory confidence labels should remain bounded. They are rule-authored mapping labels, not runtime model outputs and not empirical proof of legal acceptability.
Those limits are not weaknesses.
They are part of the trust boundary.
Try it yourself
STEM BIO-AI is Apache 2.0 and fully open source.
If you want to know whether a bio/medical AI repository is actually exposing reviewable evidence, or whether your own repository is weaker than you think, run it yourself.
That is the real test.
Final thought
The earlier STEM-AI posts were about why repository trust deserves its own audit layer.
This phase was about something more practical:
what does that audit layer have to look like if an engineer is actually going to run it, inspect it, and put it in a pipeline?
For me, the answer was simple:
Separate the workflows.
Separate the lanes.
Keep diagnostics evidence-first.
Keep regulatory mapping subordinate to evidence.
Keep advisory AI bounded.
Optimize for inspectability, not just score production.
That is what changed the project.
Not bigger claims.
Better boundaries.
