Drug discovery is expensive for a reason.
A new medicine can take roughly a decade and enormous capital to move from idea to patient. That is not simply because biology is complex. It is because biology punishes unverified claims. Every stage adds cost, but it also adds filtering. Weak hypotheses are supposed to fail early. Unsafe assumptions are supposed to be caught before they travel downstream.
That is why AI in biology is so attractive.
If models can surface candidate molecules faster, detect patterns in genomic data earlier, or automate parts of scientific analysis that used to take weeks, the upside is obvious. The promise is real. Faster iteration matters.
But over the past year, another layer has formed around that promise.
A visible class of open-source repositories now presents Bio-AI through agents, skills, wrappers, workflow kits, and automated research surfaces. They look increasingly operational. They install cleanly. They return outputs. They often present themselves in the language of discovery.
That was the background to the question I wanted to answer.
Not: Which repositories look the smartest?
Not even: Which ones are scientifically correct?
A prior question comes first.
Which of these systems provide enough structural honesty, bounded
behavior, and verification surface that someone downstream could begin
to trust what they are doing?
So I audited ten visible repositories and adjacent scientific automation systems in the open Bio-AI ecosystem.
What I found was not that most of them were obviously fraudulent, or even obviously wrong. The deeper problem was narrower and more important:
most of them had outputs, but lacked reliable mechanisms to establish
what those outputs meant, when they should stop, or how another party
would review them before acting on them.
That is not just a tooling gap.
It is a trust gap.
The Function That Changed the Entire Audit

One of the clearest examples came from a repository called OpenClaw-Medical-Skills, presented as part of a drug-discovery workflow.
def generate_analogues(self, seed_smiles: str, count: int = 3):
“”“
Mocks a generative model (like REINVENT).
In a real app, this would call a PyTorch model.
“”“
analogues = []
for i in range(count):
if “C” in seed_smiles:
new_smi = seed_smiles.replace(”C”, “C(C)”, 1) if i == 0 else seed_smiles + “F”
analogues.append(new_smi)
else:
analogues.append(seed_smiles + “C”)
return analogues
If you work in chemistry, the issue is immediate.
If you do not, the short version is simple: SMILES is not arbitrary text. It is a structured notation for molecular representations, with grammar, branching rules, ring closures, charge handling, and chemical constraints. This function does not generate meaningful analogues. It appends characters to a string.
The comment admits it is a mock. But that truth lives inside the code comment, not at the trust surface most users or downstream agents would rely on. At the level of the workflow, the repository still appears to generate candidate molecules. The function runs. The pipeline continues. A result is returned.
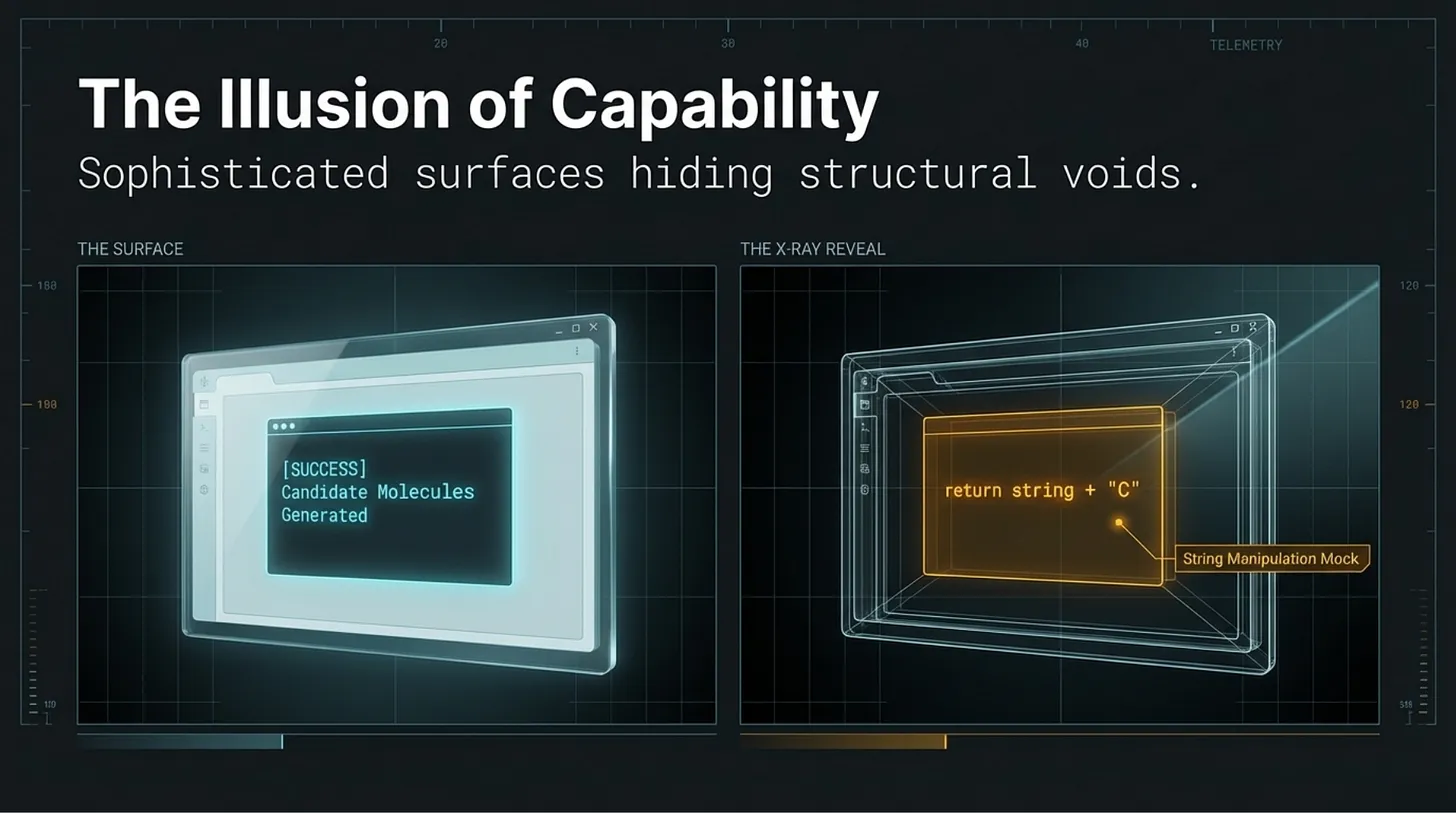

By the time the system says “candidate molecules generated,” the operational distinction between mock behavior and scientific behavior has already been blurred.
That is why I do not think this is best described as a bug.
It is a trust-surface failure.
And once I saw that clearly, I started finding related versions of the same problem across the ecosystem.
What I Audited
I reviewed ten visible repositories and adjacent systems using a two-layer process:
- a repository audit focused on structure, execution paths, default behavior, and file-level findings
- a trust-scoring framework, STEM-AI v1.0.4, focused on documentation integrity, governance posture, and biological accountability
The repositories were:
- Biomni
- AI-Scientist
- CellAgent
- ClawBio
- LabClaw
- claude-scientific-skills
- SciAgent-Skills
- BioAgents
- BioClaw
- OpenClaw-Medical-Skills
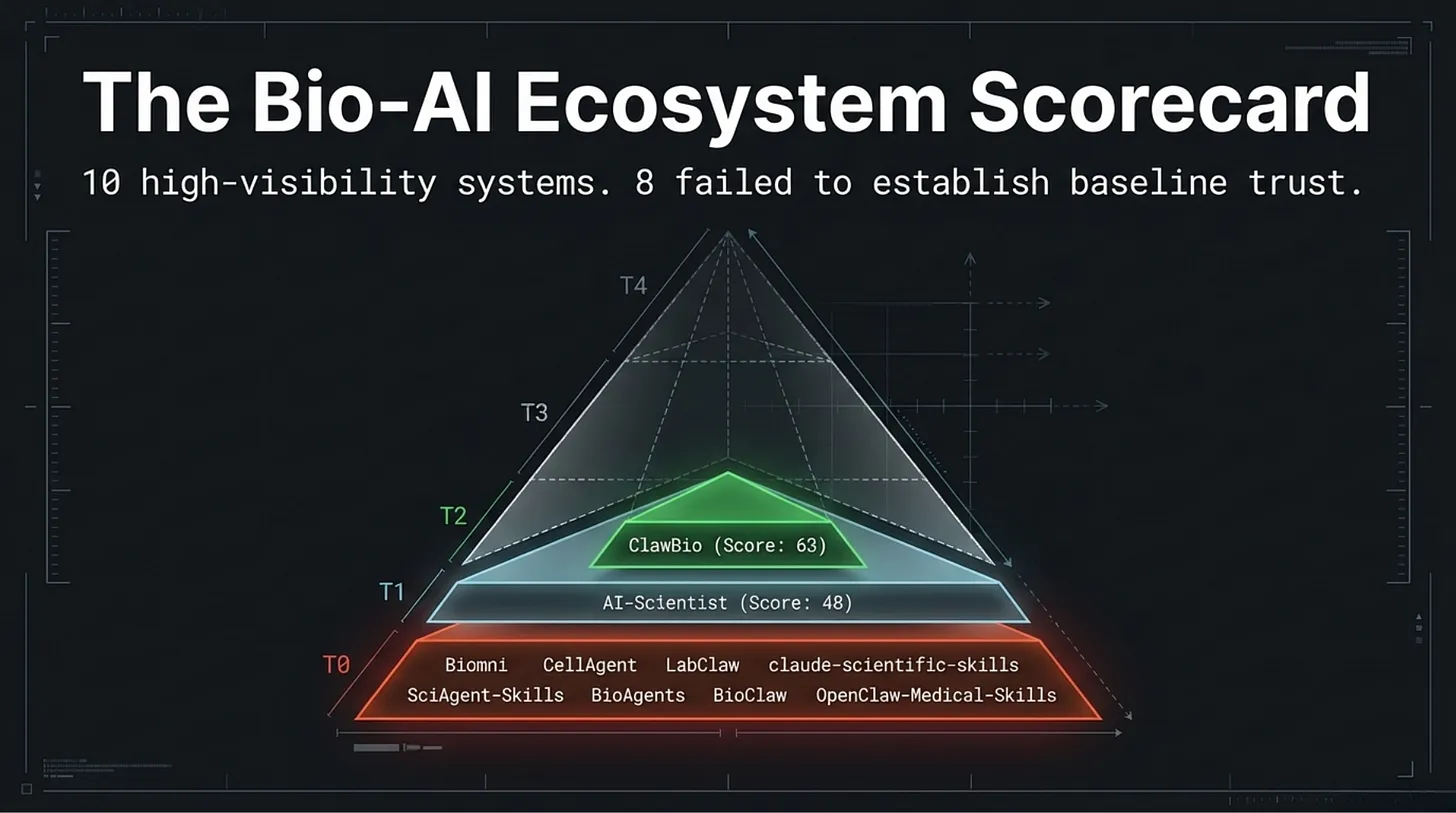

The score table looked like this:
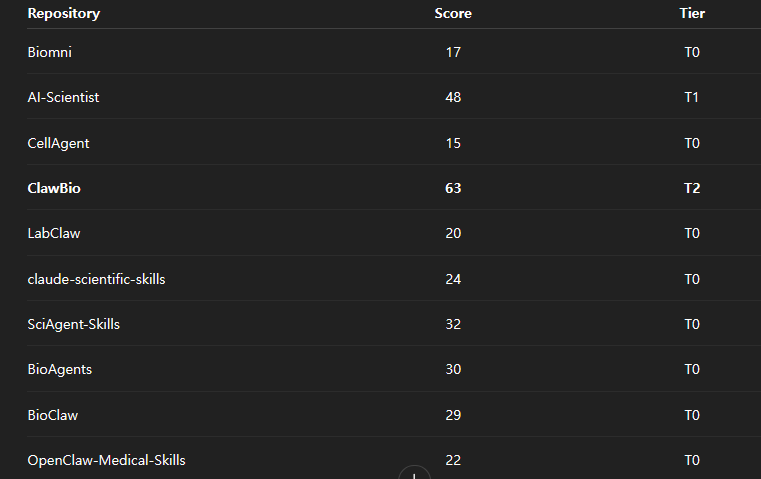
The headline result was blunt.
- 8 of 10 landed in T0
- 1 landed in T1
- 1 landed in T2
- 0 reached T3 or T4
That matters because, in this framework, T3 is the minimum threshold for supervised pilot consideration.
None of the repositories reached it.
What This Audit Was Actually Measuring

This point matters because the article can be misread if the frame is too loose.
The audit was not trying to prove which repositories were scientifically true in the strongest sense. It was asking a prior and narrower question:
Does this repository provide enough structural evidence, boundedness,
and reviewability that a researcher, team, or institution could even
begin to establish trust before downstream use?
That means asking questions such as:
- Does the README honestly describe what the system can and cannot do?
- Do critical paths fail closed when required evidence, assets, or dependencies are missing?
- Are there domain tests, not just software hygiene tests?
- Can an output be traced back to a concrete input and execution state?
- Are placeholders, stubs, or mock routines clearly separated from functional output surfaces?
That is a narrower standard than scientific truth.
But in practice it comes first.
If a system cannot establish what it did, then no downstream user can responsibly decide what its output should mean.
Four Patterns Kept Reappearing

After working through the repositories, the same four failure modes appeared again and again.
1. Scientific Scope Expanded Faster Than Accountability
Many of the repositories touched domains that are not trivial: drug discovery, molecular analysis, genomics, clinical-adjacent interpretation, or biologically consequential workflow automation.
But the surrounding accountability structure often lagged far behind the ambition of the task.
The closer a system gets to outputs that influence costly, safety-relevant, or clinically adjacent decisions, the more necessary it becomes to define limits, disclaimers, stop conditions, review boundaries, and responsibility surfaces.
Those were often weak, partial, or absent.
Several repositories had CI/CD pipelines, which sounds reassuring until you inspect what they actually verify.
In most cases, the checks focused on formatting, schema shape, ordering, script completion, or basic code hygiene. Those things matter. But they do not answer the question that matters in Bio-AI: whether the output is scientifically meaningful, biologically plausible, or safe to interpret as evidence.
Passing CI often meant the software surface was consistent.
It did not mean the scientific surface was trustworthy.
3. Mock Behavior Could Survive as Functional Behavior
The OpenClaw example was the clearest case, but not the only one.
Across the sample, I repeatedly found a dangerous pattern:
- the name of the function sounded real
- the workflow looked legitimate
- the output format appeared plausible
- but the underlying behavior was placeholder logic, a simplified stub, or a mock presented too close to a user-facing surface
Prototype code is not the problem by itself. Early repositories mock things all the time.
The problem begins when the architecture no longer makes that provisional status obvious downstream.
At that point, scaffolding can start masquerading as capability.
4. Strong Design Was Often Undermined by Weak Defaults
Several repositories were not superficially sloppy. Some had serious architecture. Some clearly reflected thoughtful engineering.
But the defaults weakened the trust story.
BioAgents presented a substantial multi-agent design, yet rate limiting could fall away in default usage. BioClaw used container isolation, but an important writable mount weakened containment. Biomni wrapped parts of execution in timeout logic, yet still exposed unsandboxed subprocess behavior.
This produced a recurring pattern I now think is characteristic of immature scientific infrastructure:
the architecture says the right things, but the runtime defaults still carry the old risks.
The One Repository That Broke the Pattern
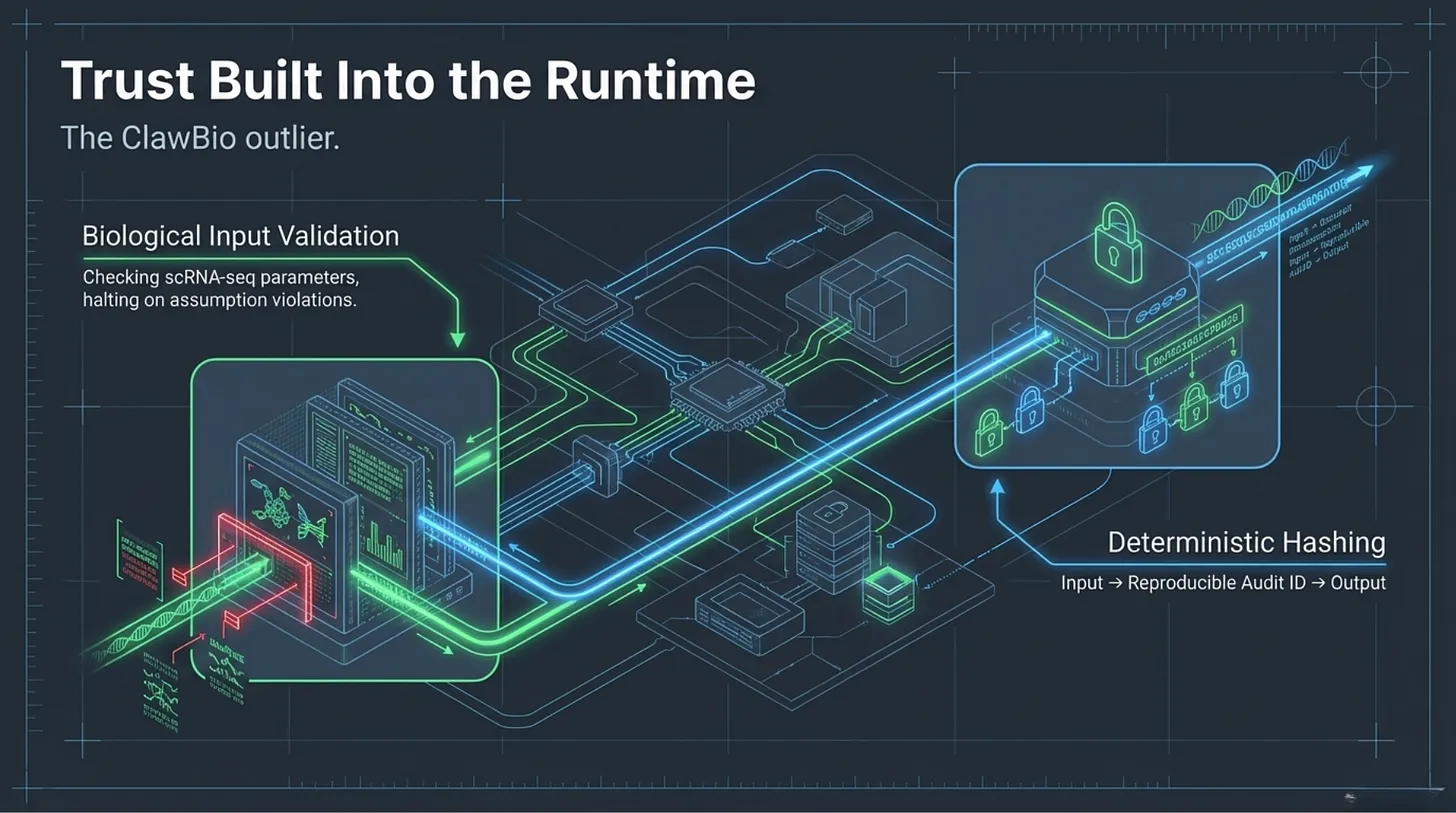
The only real outlier in the sample was ClawBio.
It did not solve everything. It still landed at T2, not T3 or T4. But it was the only repository in the set that consistently treated trust as something that had to be constructed in the runtime itself.
Two signals stood out.
First, it validated biological inputs before moving forward. Its scRNA-seq path did not simply accept a file and continue. It checked whether the data looked already processed, whether value assumptions were broken, and whether the analysis should halt rather than continue under uncertain conditions.
Second, it created reproducible audit identifiers by hashing input files. That sounds mundane. It is not. It creates a deterministic link between a specific input state and a specific output trace — exactly the kind of provenance discipline that was missing almost everywhere else in the sample.
ClawBio did not prove that open-source Bio-AI is solved.
It did prove something narrower and more useful:
the ecosystem does not have to remain trapped at the level of plausible-looking but weakly governed execution.
What Changed in My View
Before this audit, the intuitive question was:
Which Bio-AI repositories are good?
After the audit, I think that is the wrong first question.
The better first question is:
What has to exist before any Bio-AI repository can be treated as trustworthy scientific infrastructure?
That shift matters because it moves the conversation away from surface cleverness and toward institutional reviewability.
- From capability to boundedness.
- From output to provenance.
- From demo quality to stop conditions.
- From architectural ambition to governance reality.
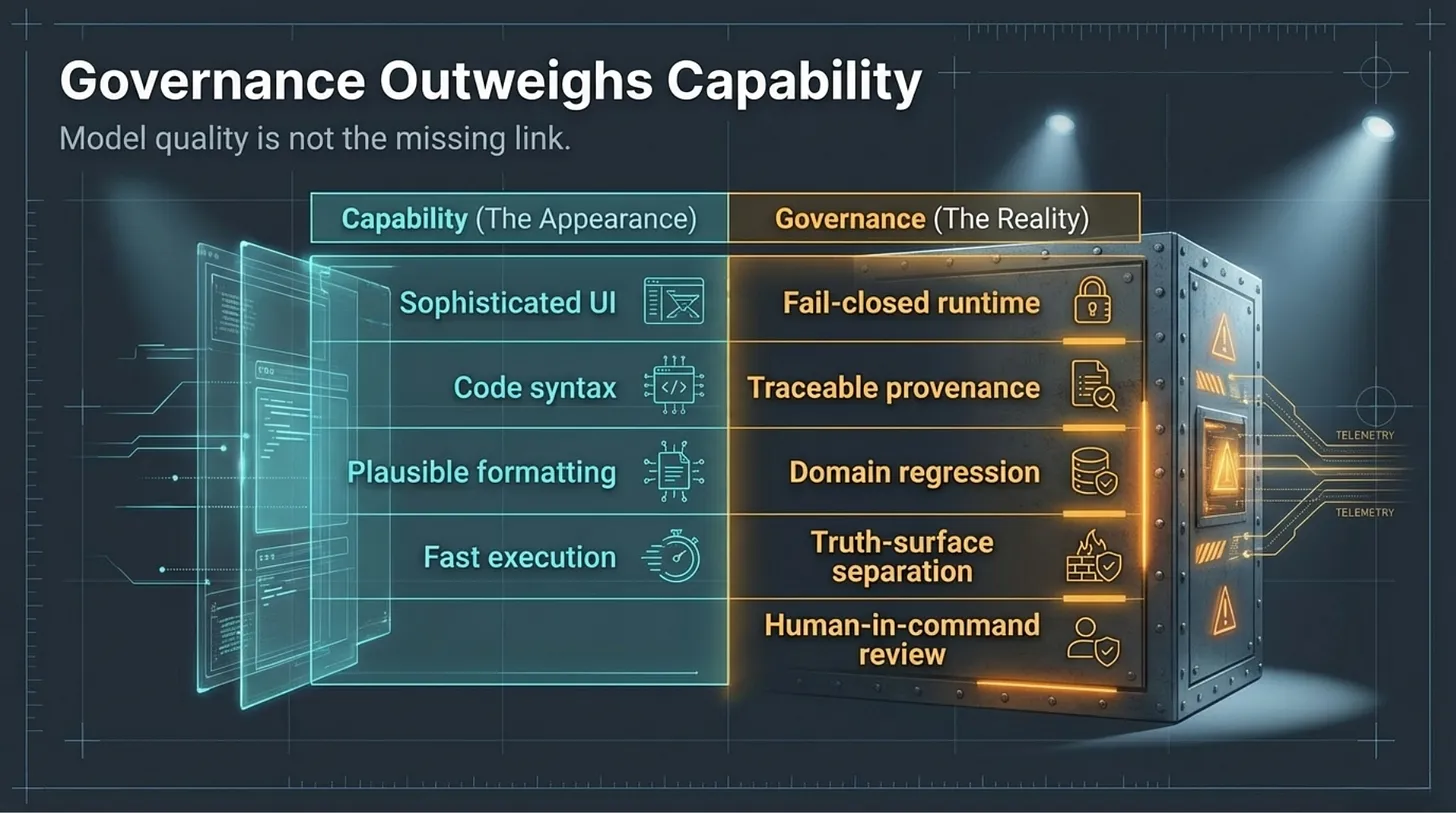
That is why I do not think the core bottleneck in Bio-AI is model capability alone.
The deeper bottleneck is the missing verification layer.
More specifically:
- truth-surface separation
- fail-closed runtime behavior
- domain regression testing
- provenance discipline
- explicit scope boundaries
- human-in-command reviewability
Without those, even technically impressive repositories remain unstable objects: too promising to dismiss, too weakly governed to trust.
The Minimum Standard I Would Use

If I were evaluating these systems for institutional use, I would not begin with benchmark claims or abstract model quality.
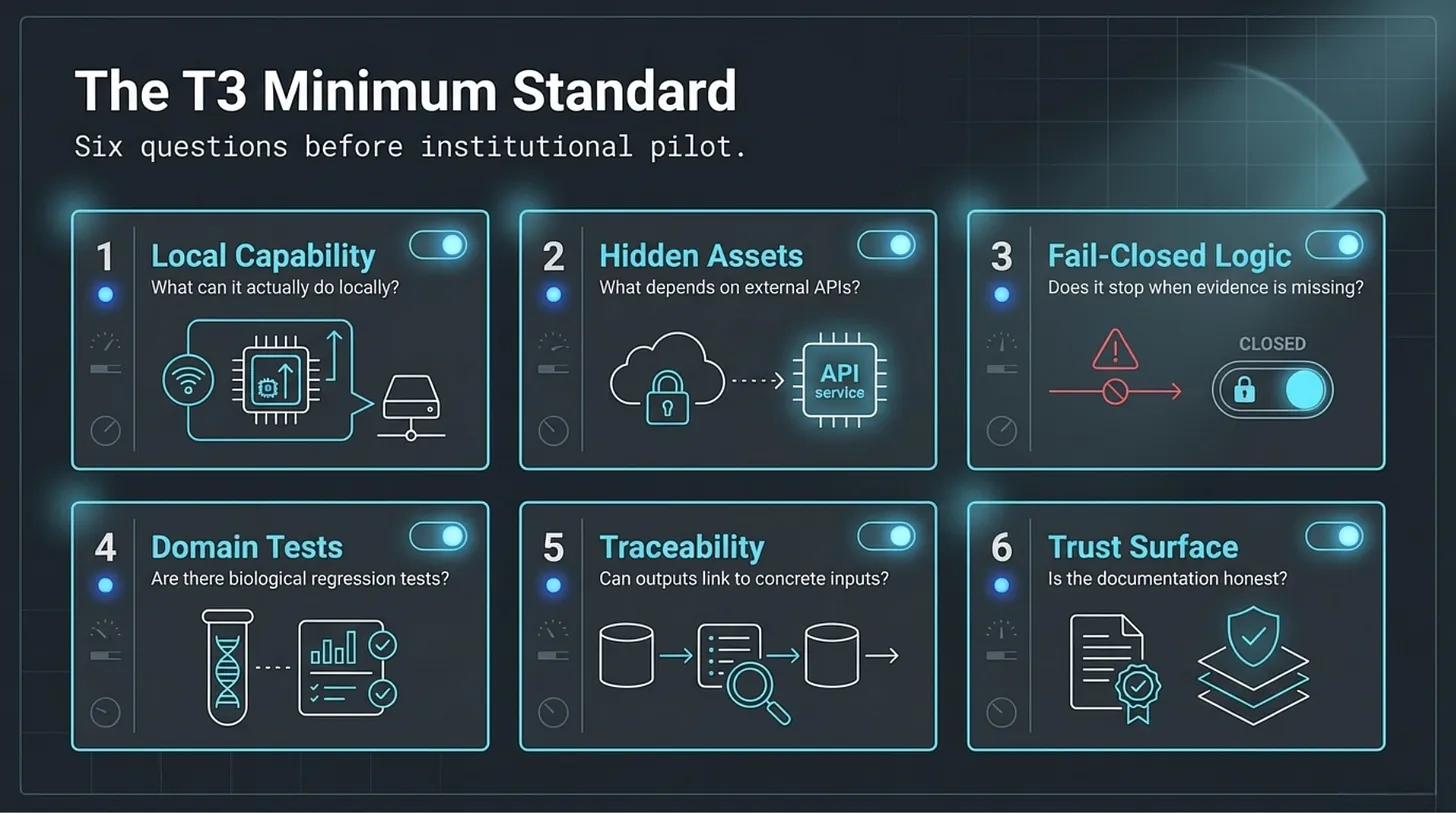
I would begin with six simpler questions:
- What can this repository actually do locally?
- What depends on hidden assets, hosted services, or private APIs?
- Does it stop when critical evidence is missing?
- Does it include domain-specific tests?
- Can outputs be traced to concrete inputs and execution paths?
- Is the trust surface honest?
Those are not glamorous questions.
They are earlier questions.
And in this audit, they mattered more than almost anything the READMEs claimed.
My own threshold would be clear:
T3 is the minimum standard for a supervised pilot.
None of the repositories in this audit reached it.
That is not a reason to stop building.
It is a reason to stop pretending that plausible output is already equivalent to trustworthy infrastructure.
Final Point
I do think AI will matter in biology.
The promise is not fake. The acceleration is not imaginary. Some of these repositories are useful research artifacts, engineering accelerators, or idea surfaces.
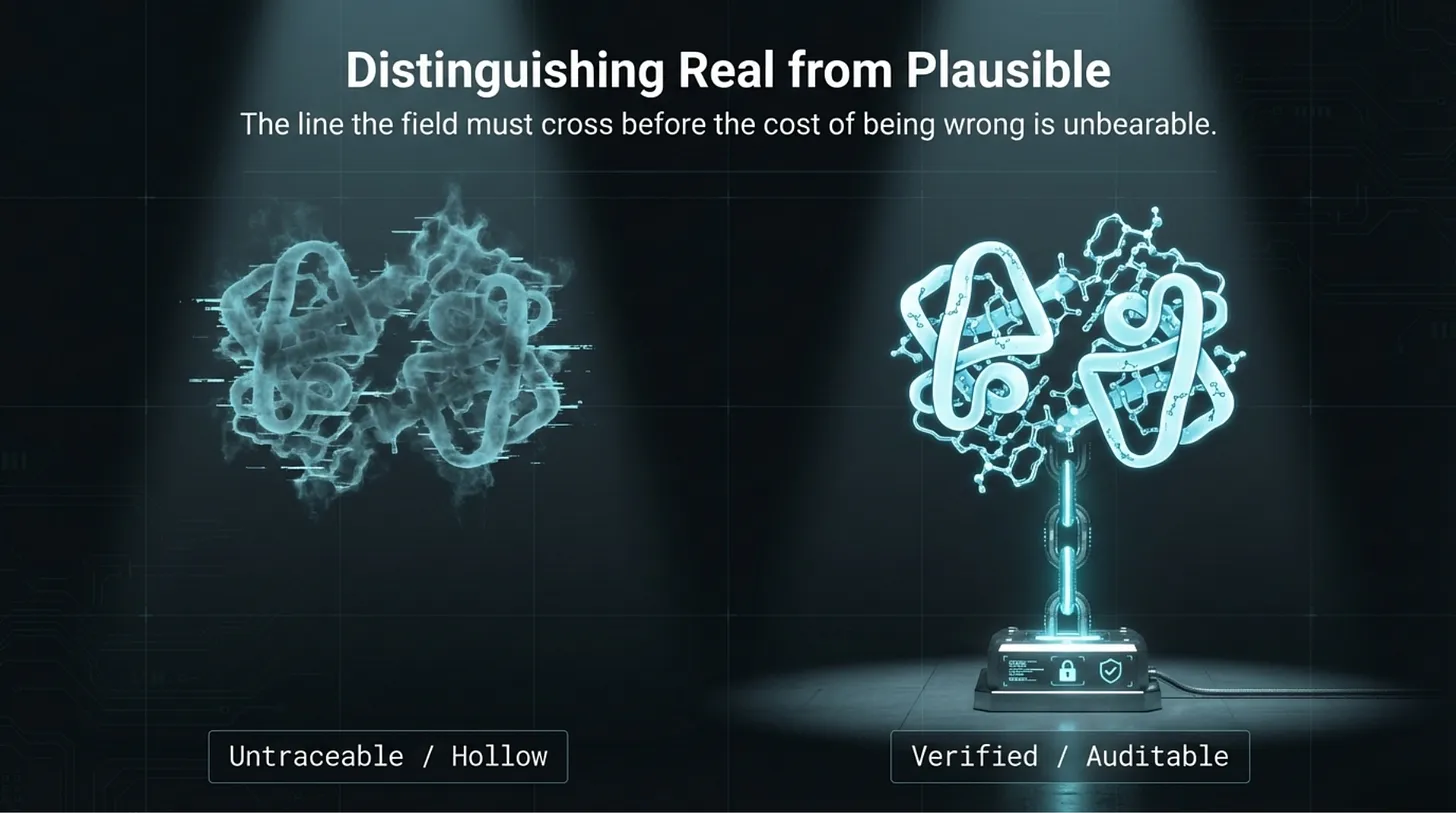
But the field is still crossing a deeper line than many people admit.
It is not just trying to produce outputs.
It is trying to produce outputs that another party can review, bound, reproduce, and challenge before those outputs acquire scientific or operational authority.
That is the line that still matters most.
And right now, in open Bio-AI, the verification layer is still lagging behind the capability layer.
Bio-AI Repository Audit 2026 — Technical Report
A technical audit of 10 open-source Bio-AI repositories using code inspection and STEM-AI trust scoring.
Audit snapshot date: March 20, 2026. STEM-AI v1.0.4.
This report reflects a time-bounded audit of public repository surfaces, workflow reconstruction, and selective file-level review.
It is not a regulatory determination or legal judgment.
→ Read the Full Technical Report
https://flamehaven.space/writing/bio-ai-repository-audit-2026-a-technical-report-on-10-open-source-systems/
Audit snapshot date: March 20, 2026. STEM-AI v1.0.4. This article reflects a time-bounded audit of public repository surfaces, workflow reconstruction, and selective file-level review. It is not a regulatory determination or legal judgment.